Three LTR-retrotransposon families compose 60% of the O. australiensis genome
In order to investigate the cause of the genomic expansion in O. australiensis, we first applied Representational Difference Analysis (RDA). RDA (Lisitsyn et al. 1993; Panaud et al. 2002) is a PCR-based cloning procedure that allows isolation of sequences which are specific to one genome (the tester) compared with another (the blocker). It is based on subtractive hybridization of genomic fractions (the representation) of both the tester and the blocker. These representations are obtained after digestion of total genomic DNA, followed by ligation with adapters and PCR amplification of the ligated products using a primer homologous to the adapter. Prior to the subtraction, a new set of adapters is ligated to the representation of the tester DNA only, thus allowing the amplification of specific sequences. Using O. australiensis genomic DNA as tester and O. sativa as blocker we obtained a library primarily composed of a single 359-bp O. australiensis-specific fragment (data not shown). This suggested that, at least in the representation we obtained, the difference in composition between these two genomes could be explained by the presence of a sequence which is highly repeated in the genome of O. australiensis but which is absent (or present at a much lower copy number) from the O. sativa genome. Sequencing of this RDA sequence revealed that it is part of the LTR of RIRE1, a previously characterized TY1/Copia type LTR-retrotransposon (Uozu et al. 1997). In order to estimate the copy number of RIRE1, dot-blot assays were performed using either LTR or internal region probes (Supplemental data #1). We found that there are 30,000 ± 3000 complete RIRE1 copies and 10,000 ± 1000 apparent single LTRs, considered as recombinational variants called solo-LTRs (Shirasu et al. 2000), which makes this element one of the most highly repeated within a plant genome. RIRE1 therefore appears to contribute a total of about 265 Mbp, i.e., 27% of the genome of O. australiensis (Table 1). This observation led us to use another strategy to identify other repeated sequences that could have contributed to the genomic expansion of the species in addition to RIRE1.
Table 1.
Description of the three retrotransposons, RIRE1, Kangourou, and Wallabi, in the genome of O. australiensis
The Oryza Map Alignment Project (OMAP, http://www.omap.org) has generated a large amount of genomic resources for 12 Oryza species, including 137,000 BAC end sequences (BES) ofO. australiensis (Ammiraju et al. 2006). In silico analyses of theseO. australiensis BES allowed identification of 25 highly redundant sequences which were not homologous to RIRE1. These were successively extended and merged using the BES data, allowing reconstruction of the putative sequence of three TY3/Gypsy type LTR-retrotransposons, named Kangourou, Wallabi, and Dingo (Fig. 2; Supplemental data #2). Three BAC clones from the O. australiensis genomic libraries were then fully sequenced to validate the structure of these three new elements. One clone harbors the HD1 locus (and was chosen because it also harbors at least one copy of RIRE1). The other two clones were chosen based on the homology of one of their BES with either Wallabi or Kangourou. Comparative sequence analysis with the O. sativa genome suggests that none of the three clones are located in a pericentromeric region. Overall, 350 kb (350,792 bp) of genomic sequence were generated and analyzed (Fig. 3). Seven complete copies and five solo-LTRs of RIRE1, Kangourou, and Wallabi were identified from these genomic clones, thus validating the sequences inferred from the in silico approach. Moreover, Kangourou and Wallabi were searched for homology with already known rice retrotransposons. We found that Kangourou exhibits a low but significant sequence identity with the Retrosat1/RIRE2 retrotransposon family (65% overall identity in the internal region with 75% in the GAG-POL region). This is an indication that these two families are homologous. However, given the low sequence identity between Kangourou and Retrosat1, we kept a distinct name for each.
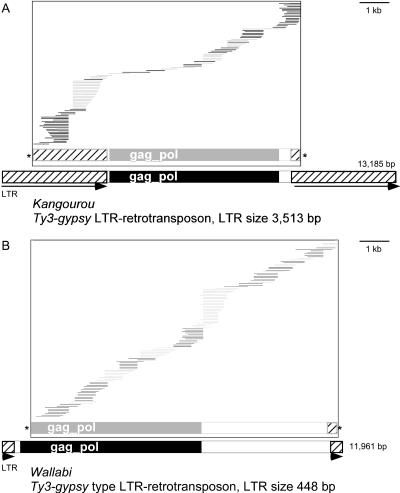
Figure 2.
In silico reconstruction of Kangourou (A) and Wallabi (B) retrotransposons. The copies of the BES contigs are shown as horizontal lines (alternate black and gray according to their final position on the element). The schematic representation of the element assembled from O. australiensis BES is represented in gray. The schematic representation of the elements, as it is found in the O. australiensis sequenced BAC clones, is given in black at the bottom of the figures. The size scale in bp is given at the bottom.
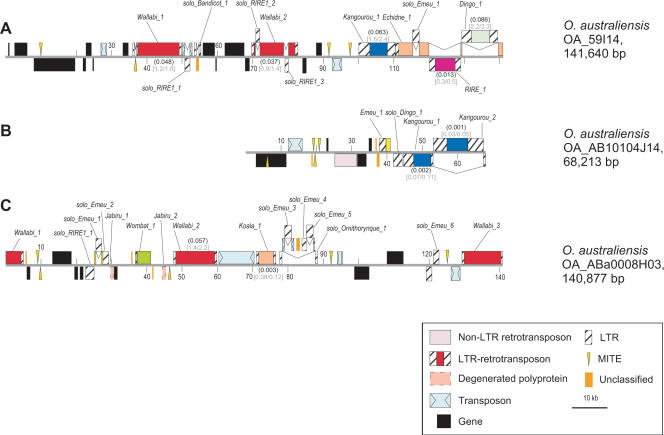
Figure 3.
Physical map of three sequenced O. australiensis BAC clones. Black boxes represent predicted coding regions. Colored boxes represent different types of TEs as indicated on the figure. Numbers in parentheses indicate the estimated date of LTR-retrotransposon insertions (in million years) using the two molecular clocks MC1 and MC2 (see text). A,B,C correspond to the sequence of the BAC clones OA_59114, OA_AB10104J14, and OA_ABa0008H03, respectively.
Based on dot-blot assays, Kangourou and Wallabi were estimated to contribute a total of 90 ± 9 Mbp and 250 ± 25 Mbp,i.e., 9% and 26% of the genome of O. australiensis, respectively (Table 1). The copy number of Dingo was estimated to be between 3700 and 4300 and could thus clearly be considered as highly repeated, but its contribution to the genomic expansion of O. australiensis (i.e., <5% of the present size) was considered negligible compared with the other three elements. Dingo was therefore not included in further analyses. Altogether, RIRE1, Kangourou, and Wallabi contribute ∼60% (605 ± 40 Mbp) of theO. australiensis genome (Table 1). In contrast, BLAST searches of the genomic sequence of O. sativa (cv. Nipponbare) revealed that it contains only two, 10, and 16 complete copies of RIRE1, Kangourou, and Wallabi elements, respectively. The retrotransposition bursts of these three elements alone could thus account for the increase in the genome size of O. australiensis, compared with that of O. sativa. Our data therefore show that, at least in the case of the genus Oryza, retrotransposition has contributed to genome size variation to an extent which is comparable with that of polyploidization.
We first anticipated that BES may not be a random representation of the genome of O. australiensis, mainly because the BAC library was constructed using the HindIII restriction enzyme, but our results show a posteriori that this approach was nevertheless successful for retrieving the most highly repeated elements from the genome regardless of the representational bias that may have been caused by the restriction enzyme. In fact, it is clear from Figure 2 that some regions of the retrotransposon are overrepresented in the BES database, probably because most of the paralogs harbor a HindIII site at this location. Contrastingly, it is expected that BES corresponding to the regions of the element where the majority of the paralogs do not harbor a cleavage site should be far less frequent. However, in our case, we were able to retrieve at least one BES in all the regions of the three elements, probably because of the high copy number of each family, thus allowing their in silico reconstruction.
Deep annotation of the sequences of the three BAC clones revealed the presence of many TEs which are distinct from RIRE1, Kangourou, and Wallabi (Fig. 3). In all, 61 putative transposable elements were identified, accounting for 195,277 bp of sequence and representing 55.6% of the complete BAC sequences. Among these, a majority belong to the Class I retrotransposon group (accounting for 65% of all the TE and 42.9% of the BAC sequences). The 39 LTR-retrotransposons identified in these sequences belong to 11 distinct families. The LTR-retrotransposons, analyzed and annotated in the three sequenced BAC clones, were found mainly clustered within intergenic regions (Fig. 3). These elements are frequently disrupted by successive insertions of other LTR-retrotransposons, leading to the formation of nested structures as often observed in larger genomes of other cereal species (SanMiguel et al. 1996; Wicker et al. 2001, 2003). Distal parts of BACs OA_59I14 and OA_AB10104J14 and the central part of BAC OA_ABa0008H03 showed a high density of clustered and nested LTR-retrotransposons, with respectively five, four, and eight elements clustered within a distance of 41, 30, and 50 kb. Altogether, Kangourou, Wallabi, and RIRE1 account for ∼29% of all the BAC sequences which is much less than their overall genome contribution (60%). However, FISH experiments previously revealed that RIRE1 elements are more abundant in pericentromeric than in distal regions of O. australiensis chromosome arms (Uozu et al. 1997), a characteristic shared with other LTR-retrotransposon families in theO. sativa genome (Jiang et al. 2002; Vitte and Panaud 2003). Consequently, the distribution of the elements in the three BAC sequences may not reflect their actual distribution in the genome.
The genomic amplification of O. australiensis occurred after its speciation
In order to trace the origin of the three elements RIRE1, Kangourou, and Wallabi, we surveyed their presence in nine different genome types of the genus Oryza by Southern hybridization using probes corresponding to either the LTR or the internal region (Fig. 4). The results clearly show that all three elements are present in at least one other wild Oryza species, indicating an ancient origin in the genus. Moreover, the strong hybridization signals obtained for some species distantly related to O. australiensis (e.g., in O. granulata [GG] for Wallabi) suggest that independent transposition bursts of RIRE1 and Wallabi have occurred in distinct genome types of the genus, although to a lesser extent than in the genome of O. australiensis. In order to tentatively characterize and date the transposition bursts of the three elements in the O. australiensis genome, we conducted phenetic analyses of the three elements based on the OMAP BES data of the 12 Oryza species (Fig. 5; Supplemental data #3): For all three elements, the paralogs found in O. australiensis form a cluster which is distinct from those found in other Oryza species (i.e., supported by a bootstrap value which is >70%), suggesting that the retrotransposition bursts occurred concomitantly or after the speciation of O. australiensis.
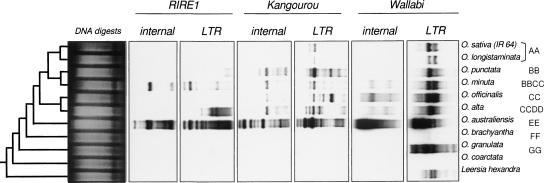
Figure 4.
Southern hybridization of the three retrotransposons, RIRE1, Kangourou, and Wallabi on total genomic DNA of Oryza species digested with RsaI. The phylogenetic tree given on the figure is extrapolated from Ge et al. (1999). The direction of migration is from left to right.
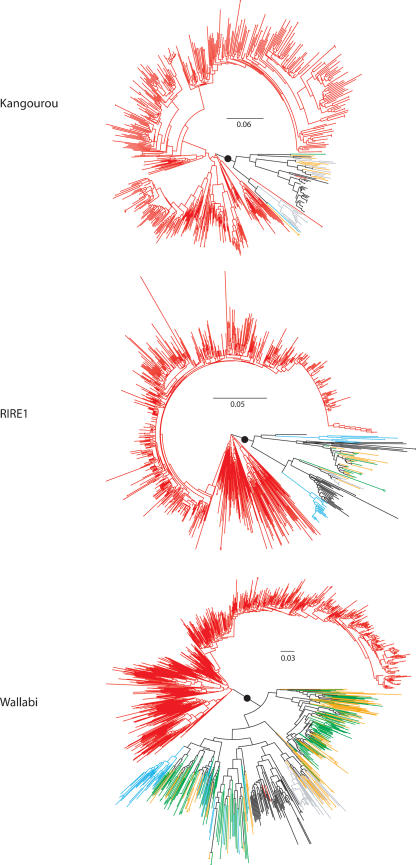
Figure 5.
Phenetic relationships of RIRE1, Kangourou, and Wallabi in the genus Oryza: The neighbor-joining tree was constructed based on the alignments given in Supplemental data #3. For each tree, the dot shows the branch separating the O. australiensis sequences from the others. The number given near the dot corresponds to the bootstrap value. Color coding: black for the A-genome species; gray for O. punctata; orange forO. minuta; green for O. officinalis; blue for O. alta, and red for O. australiensis. The numbers of aligned sequences used to build the tree were as follows: for RIRE1: 752 O. australiensis sequences and 113 other Oryza sequences; for Kangourou: 570 O. australiensis sequences and 67 others; for Wallabi: 757 O. australiensis sequences and 422 others.
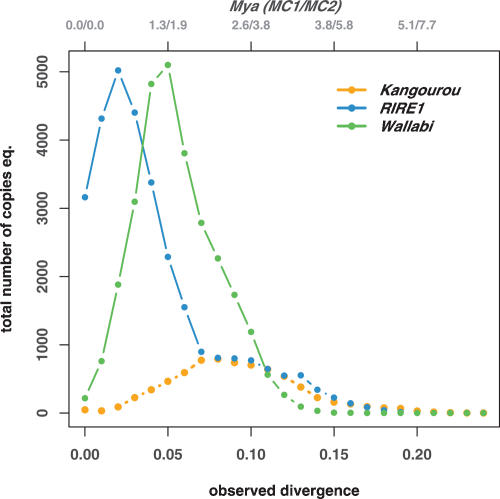
In order to test this hypothesis, the dates of retrotransposition of RIRE1, Kangourou, and Wallabi were estimated in the O. australiensis genome (Fig. 6). We applied an approach of genomic paleontology, which consists of translating the nucleotide divergence observed between the paralogs mined out from the BES into a radiation date. This approach relies on the estimation of the rate of the molecular clock (MC) of retrotransposon sequences once they are inserted in the genome. The first examples of such studies in plants were conducted using an MC of6.5 × 10−9 synonymous substitutions/site/year (SanMiguel et al. 1998), an estimation based on the MC of the ADH2 gene in the Poaceae family (Gaut et al. 1996). Several subsequent studies have led to a re-estimation of the MC of retrotransposons in rice, i.e., 2 × 10−8 subst/site/year (Vitte et al. 2004), referred to as MC1, and 1.3 × 10−8 subst/site/year (Ma et al. 2004), referred to as MC2. The data provided in the present paper are given using both these new MC. In any case, the translated dates can only be considered as rough estimates and only large differences should be retained as putatively significant. The figure clearly shows that the transpositional activity of the three elements has not been continuous during the last 3 to 4 Myr: A peak of activity (defined here as a burst) is indeed observed at ∼0.5–0.75, 1.2–1.8, and 2–3 Mya for RIRE1, Wallabi, and Kangourou, respectively. Moreover, the size of the peaks shown in Figure 6 is proportional to the number of complete copies of the corresponding elements in theO. australiensis genome. This representation clearly shows that the largest bursts are the most recent and, therefore, that most of the genomic expansions that led to the doubling of the genome size of O. australiensis are of recent origin (i.e., within the last 3 or 4 Myr). This is further supported by dating of the insertions of RIRE1, Kangourou, and Wallabi elements found in the three BAC sequences (Fig. 3). In the case of full-length LTR-retrotransposons, the date of insertion can be estimated based on the divergence between their two LTRs (SanMiguel et al. 1998; Vitte et al. 2004). The estimated insertion time of these elements ranges from 1.6 to 2.4 Mya for Kangourou_1 (OA_59I14) to 0.03–0.05 Mya for Kangourou_2 (OA_AB10104J14). Because the date of the radiation of O. australiensis species is estimated at 8.5 Myr (Fig. 1), we conclude from all these lines of evidence that the genomic expansion is posterior to and not concomitant with the speciation. These results also suggest that the strong hybridization signals observed in some other Oryza species in the Southern hybridization experiments (e.g., for Wallabi in O. granulata, Fig. 4) reflect distinct bursts of the corresponding elements in these lineages. In this regard, the large size of the O. granulata genome(i.e., 880 Mbp) (Ammiraju et al. 2006), compared with that ofO. sativa, may be partly accounted for by the retrotranspositional activity of Wallabi, although more detailed analyses are needed to quantify precisely this contribution.
Figure 6.
Timing of the bursts of the three retrotransposons, RIRE1, Kangourou, and Wallabi: For each element, the curves represent the distribution of the observed divergence between each paralog (given at the bottom x-axis). Top x-axis represents the date of divergence in Mya translated from the observed divergence, using the two molecular clocks MC1 and MC2 (see Methods section). The groups of paralogs used to compute the pairwise distances are defined within the phenetic subgroups shown in the phenogrammes given in Supplemental data #4. The y-axis represents the total number of copy equivalent, i.e., (the frequency at which the divergence time occurred) × (the number of paralogs in the genome of O. australiensis, based on the dot-blot experiments, Table 1).
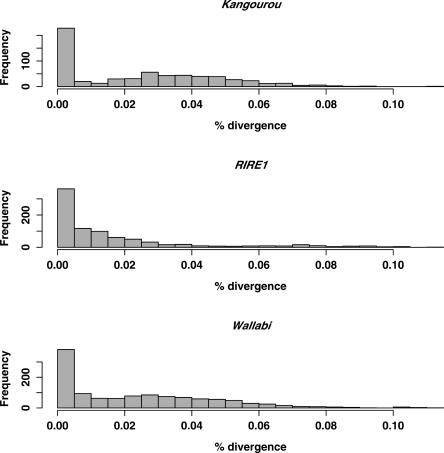
The phenetic analyses also provide interesting insights into the dynamics of the genomic expansions observed in O. australiensis. The peaks shown in Figure 6 are not overlapping, thus suggesting that the maximum transpositional activity of the three elements did not occur concomitantly. The cause of these successive waves of retrotransposition could be regulatory, i.e., the result of an activation (triggered by external stimuli, such as biotic or abiotic stress) and/or by the repression of silencing of the corresponding elements. Alternatively, these distinct bursts may be explained by the presence of active elements in the genome only during a short period (corresponding to the peaks). These active elements may have arisen from ectopic recombinations between two defective copies, a mechanism known for retroviruses (Bartosch et al. 2004). The presence of RIRE1, Kangourou, and Wallabi in the genome of many other Oryza species (Fig. 4) suggests, however, that functional copies of these three elements have probably been present in the genus since its origin. In order to test whether active copies may still be present in the genome of O. australiensis, we determined for each paralog the shortest distance found among all the pairwise distances computed with all the other copies (at the nucleotide level). The distributions of these shortest distances are given in Figure 7. Interestingly, for RIRE1, Kangourou, and Wallabi, several pairs of very closely related paralogs can be found (the first bar of the histogram), suggesting recent transposition (<200,000 yr ago) of active elements. Consequently, transpositional bursts observed in this species may have their origin in a regulatory process, rather than a structural mechanism, although this remains a hypothesis that should be further tested. As a first step one should conduct expression studies of the three elements in O. australiensis in order to assess whether its genome still harbors transcriptionally active copies.
Figure 7.
Histograms of the most recent among all observed divergence computed for each paralog (compared with all others) of the three retrotransposons RIRE1, Kangourou, and Wallabi.
Several reports on both plants and animals have shown that transposable elements are efficiently eliminated from eukaryotic genomes, either by recombination or deletion (Petrov et al. 1996; Shirasu et al. 2000; Ma et al. 2004; Chantret et al. 2005). In particular, LTR-retrotransposons tend to be partially eliminated through ectopic recombinations between their two LTRs, leading to the formation of solo-LTRs (Shirasu et al. 2000). This was taken into account in our estimation of the total contribution of the three elements to the genome size increase of O. australiensis by using both LTR and internal region probes in the dot-blot assay (Table 1; Supplemental data #1). Interestingly, the apparent single LTRs represent a significant percentage of the total number of copies that we estimated (i.e., 25%, 9.5%, and 30% for RIRE1, Kangourou, and Wallabi, respectively), regardless of the age of the bursts (that of RIRE1 being the most recent). This corroborates earlier reports suggesting that the process of partial removal of LTR-retrotransposons through ectopic recombinations leading to solo-LTRs is concomitant with (or occurs shortly after) retrotrans- position (Vitte and Panaud 2003). Illegitimate recombination mechanisms targeting LTR-retrotransposons have been identified as inducing considerable loss of DNA and contributing to genome size reduction in Arabidopsis and rice (Petrov 2002; Ma et al. 2004). Analysis by sequence alignment of Wallabi, Kangourou, and RIRE1 LTR-retrotransposon families from the three fully sequenced BAC clones (Fig. 3) revealed the presence of limited small deletions, corresponding to 2.4%, 6.7%, and 2.6% of the total length of the Wallabi, Kangourou, and RIRE1 elements, respectively (data not shown). Altogether, these results indicate that processes leading to the elimination of retrotransposon sequences such as unequal and illegitimate recombinations occurred in the genome of O. australiensis similarly to O. sativa. Both the extent and timing of these DNA losses need to be investigated in order to clarify their overall contribution to the O. australiensis genome size variation following the bursts of the three retrotransposon families. Nevertheless, we anticipate that the overall DNA loss of the three LTR-retrotransposons through small illegitimate recombinations is negligible and did not lead to overestimation of their overall contribution in the genome of the species, based on our dot-blot assay.
The current model of eukaryotic genome evolution in relation to the activity of TEs posits that genome size should result from two balanced forces: increase, induced by retrotransposition, and decrease, caused by recombinations and deletions (Petrov 2002; Vitte and Panaud 2005). The evolutionary dynamics of RIRE1, Kangourou, and Wallabi in the genus Oryza provides an opportunity to test this hypothesis. Our Southern hybridization and phenetic data suggest that the three elements were present in the genome of the ancestor of the genus. This is further supported by the presence of homologs of these elements in BES of nearly all the Oryza species of the OMAP project. We also show that they have undergone independent amplification in distinct lineages in the genus, leading to one case of genomic obesity (i.e., in O. australiensis). In other lineages, their strong regulation and elimination may have led to a decrease in genome size (e.g., in O. glaberrima), but this still remains purely speculative. In this regard one should point out that, if all the copies of the three retrotransposons families RIRE1, Kangourou, and Wallabi were to be removed from the genome of O. australiensis, about 360 Mbp of genomic DNA would remain, i.e., a size comparable with that of the smallest diploid genomes of the Oryza genus. Further examination of complete sequences of high copy number LTR- retrotransposons in O. australiensis will provide better insights into the dynamics of the elimination process. The model also predicts that the successive events of TE insertions followed by their elimination should cause a fast turnover of intergenic regions, leading to their rapid divergence among distinct evolutionary lineages. This has been confirmed in several comparative studies between the genomes of maize, sorghum, and rice (Tikhonov et al. 1999; Ma et al. 2005). Comparative genomics studies within the Oryza genus, and in particular between closely-related species, would allow us to test this hypothesis and give insight into the dynamics of the process.
Maize, wheat, and barley are large genomes that contain 50%–80% of LTR-retrotransposons. It is now commonly accepted that retrotranspositions have played a crucial role in genomic expansion and architecture and could also have an impact on the transcriptional regulation of genes of these major crop species (Kashkush et al. 2003). However, the relatively older burst of amplification of these elements and the limited genomic sequence information from these large genomes make it difficult to reconstruct the history and study the impact of retrotransposon amplification at the whole genome level. The present study provides the first direct evidence that active LTR-retrotransposons can contribute to large variations in genome size over short periods of time, i.e., at the species level. As in the case of maize, wheat, and barley, LTR-retrotransposons are the main component of the O. australiensis genome. Cytologically, the mitotic chromosomes of O. australiensis show a twofold size increase compared with the Oryza sativa species (Uozu et al. 1997), indicating their dramatic impact on chromosome morphology and suggesting major events of genome reshaping. The availability of comprehensive genomic resources for many species in the genus Oryza makes possible physical comparison between closely related genomes contrasting for their size and will allow studies on the evolutionary history of the LTR-retrotransposons at the genus scale. This provides a unique and promising opportunity to unlock our knowledge on the causes of retrotransposition bursts as well as their impact on plant genome architecture and gene expression.