Gene regulatory elements, such as enhancers and promoters, regulate lineage- and region-specific transcription in the developing human cortex (10). Promoters are located adjacent to their target genes whereas enhancers can be located at distal locations from the gene(s) that they regulate. In addition, because of their cell-type specificity and spatiotemporal dynamic activity, enhancers are difficult to identify. Single-cell assay for transposase-accessible chromatin with sequencing (scATAC-seq) at different developmental stages of the human cortex enabled the identification of different cell populations and their candidate regulatory elements (11, 12). However, these studies are descriptive and do not provide a functional readout that can test enhancer activity and the effects of variants. Massively parallel reporter assays (MPRAs) allow quantification of enhancer activity in a high-throughput manner, including different alleles in a single experiment (13–16). Machine learning can leverage MPRAs and other genomic data to predict enhancers and their quantitative activity (17–22). Sequence-based deep learning models (23, 24) have been deployed at scale to screen variants prior to experimental validation and to design cell type–specific enhancers. These strategies shed light on the sequence motifs and upstream regulators that are important for regulating gene expression across different cell types and species.
We used deep learning and a lentivirus-based MPRA (lentiMPRA) to characterize the enhancer activity of 102,767 sequences in primary human mid-gestation cortical cells and 10-week cerebral organoids, each tested across 3 to 5 replicates. We discovered 46,802 functional enhancers and 164 variants with allelic differences in enhancer activity regulating known disorder-associated genes such as TBR1, MARK2 (ASD), and NFKB2 (schizophrenia). We observed comparable activity between organoids and primary cells, suggesting that organoids provide an adequate model to study the developing cortical regulatory landscape. Using our lentiMPRA data, we trained a deep learning model that predicts enhancer activity from sequence with state-of-the-art accuracy, enabling us to learn sequence determinants and upstream regulators of human cortical development. These findings provide a comprehensive catalog of functional cortical enhancers and variants that alter their activity, improving our understanding of the molecular basis of neurodevelopment.
Results
LentiMPRA library generation and analysis
To comprehensively characterize human neurodevelopmental enhancers and their sequence variants in the mid-gestation cortex, we designed two lentiMPRA libraries (25) and tested them in primary human cortical cells (Fig. 1A). Because of the limited number of obtainable human primary cells and lentivirus integrations into these cells, each library was assayed independently.
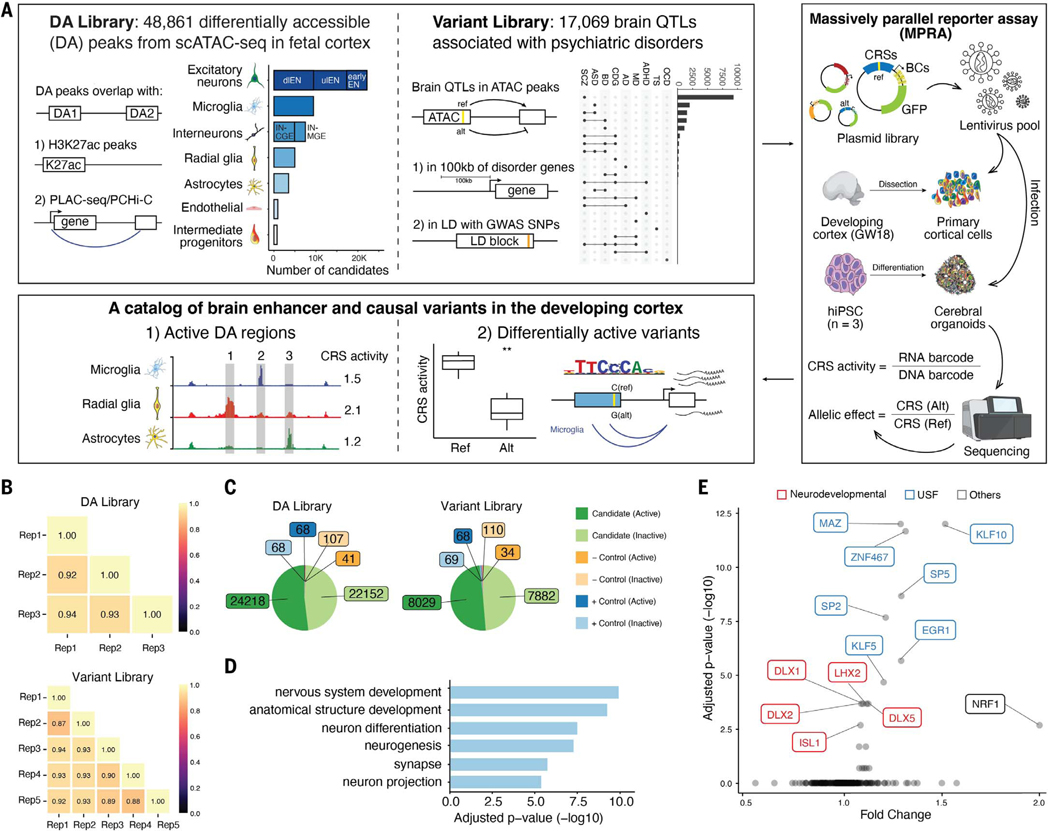
Fig. 1. Design and overall lentiMPRA results.
(A) Experimental overview of the two lentiMPRA libraries. The DA library contains 48,861 DA regions from scATAC-seq in the developing human cortex that overlap either H3K27ac peaks or PLAC-seq/PCHi-C loops. The number of DA candidates for each cell type is illustrated in the bar plot. dlEN, deep layer excitatory neuron; ulEN, upper layer excitatory neuron; IN-CGE, caudal ganglionic eminence–derived interneurons; IN-MGE, medial ganglionic eminence–derived interneurons. The variant library includes 17,069 brain QTLs that are within 100 kb of differentially expressed cross-disorder neurodevelopmental genes or in linkage disequilibrium (LD) with psychiatric disorder GWAS SNPs. The number of variants associated with each disorder is shown in the upset plot (largest 22 intersections shown). SCZ, schizophrenia; ASD, autism spectrum disorder; BD, bipolar disorder; CDG, congenital disorders of glycosylation; AD, Alzheimer’s disease; MD, major depression; ADHD, attention-deficit/hyperactivity disorder; TS, Tourette syndrome; OCD, obsessive-compulsive disorder. Both libraries were cloned into a lentiMPRA vector and packaged into lentivirus and used to infect primary cortical cells dissociated from GW18 tissues and human induced pluripotent cell (hiPSC)-derived cerebral organoids. Following infection, DNA and RNA were extracted and sequenced and an RNA/DNA barcode count ratio was calculated for each candidate regulatory sequence (CRS) allowing the identification of active DA regions and differentially active variants. (B) Correlation of log2(RNA/DNA) between technical replicates in primary cortical cells for the DA and variant library, respectively. (C) Pie charts showing the number of active and inactive sequences for candidates, positive (+) and negative (−) controls in both libraries. (D) Top enriched GO terms from the “biological process”, “ cellular component”, and “molecular function” ontologies for nearest genes of the highest activity sequences (both libraries combined). Closest genes of the lowest activity sequences were used as the background set. The complete list of GO terms is available in fig. S2B. (E) TF motif enrichment analysis for highest activity sequences (both libraries). Red, neurodevelopmental TFs, Blue, USFs.
The differentially accessible (DA) library was designed to characterize the regulatory potential of candidate cell type–specific enhancers. It consisted of 51,495 sequences obtained primarily from scATAC-seq DA peaks in the developing human brain (11). These DA peaks were further selected based on their (i) overlap with H3K27ac peaks from bulk prefrontal cortex tissue (26), microglia or non-microglia cells (11) (n = 24,611, 53%); (ii) overlap with H3K4me3 proximity ligation-assisted chromatin immunoprecipitation sequencing (PLAC-seq) peaks from intermediate progenitor cells, radial glia (RG), excitatory neurons (EN), or interneurons (IN) (27) (n = 12,412, 26.8%); or (iii) overlap with promoter capture Hi-C (PCHi-C) from EN, hippocampal dentate gyrus (GE)–like neurons, lower motor neurons and astrocytes (28) (n = 13,712, 29.5%).
The variant library compared the reference and the alternative alleles for 17,069 variants. These were selected from pseudo-bulked ATAC-seq peaks (11) overlapping brain quantitative trait loci (QTLs) (29–31). As the median enhancer-target distance was estimated at 62 kilobases (kb) (32), we further required that expression QTL (eQTL; n = 14,021) and chromatin QTL (caQTL; n =149) be within 100 kb of genes differentially expressed in schizophrenia, autism, or bipolar disorder. To overcome the systematic bias toward different types of variants in genome-wide association studies (GWAS) versus QTL studies (33), we also included QTLs in LD blocks with GWAS single nucleotide polymorphisms (SNPs) for various psychiatric disorders (8, 34–41) (eQTL n = 2,882, caQTL n = 17).
To prioritize distal enhancers, promoter-overlapping peaks were excluded from both libraries. Each library contained 143 positive control sequences nominated from ATAC-seq and ChIP-seq data in brain organoid models (42) and used to define active sequences. The variant library also included ~15,000 non-QTL sequences with a range of expected activity levels predicted from their epigenetic profiles. We designed 270 base pair (bp) oligos, each centered on the DA peak summit (DA library) or variant (variant library), flanked by 15-bp adapters on either side for library amplification. A 31-bp minimal promoter and 15-bp random barcode were placed downstream of each synthesized oligo through PCR and cloned into a lentiMPRA vector (Fig. 1A).
Each library was packaged into lentivirus and used to infect gestational week-18 (GW-18) human primary cortical cells following two days in culture. The presence of major cortical cell types was confirmed by immunocytochemistry before and after infection (fig. S1A) and single-cell RNA-seq (fig. S1B). Utilizing single-cell RNA-seq performed on infected cells, we confirmed sufficient infection rates in most cell types (fig. S1C). We performed three replicates for the DA library and five replicates for the variant library. Three days post infection, when most of the nonintegrated virus degrades, DNA and RNA were harvested and prepared for sequencing. DNA sequencing revealed that both libraries contained more than 96% of the designed oligos (DA library: 50,394 oligos; variant library: 51,319 oligos), and each oligo had an average of more than 50 unique barcode associations (median DA: 56, variant: 64). Overall, 97,762 sequences (95%) passed stringent quality control (25).
To measure enhancer activity, we quantified depth-normalized barcode abundance in DNA and RNA for each oligo and then calculated its batch-corrected RNA/DNA ratio, observing sufficient reproducibility (average Pearson correlation between replicates, DA: 0.93, variant: 0.91; Fig. 1B). We next compared the activity distributions of positive and negative controls (fig. S2A). As expected, positive controls had significantly higher ratios than negative controls (DA: P = 1 × 10−3, variant: P = 8 × 10−5, Wilcoxon test). Moreover, the distribution of ratios for randomly scrambled controls was highly comparable between libraries (median DA: 0.997, median variant: 0.994).
To identify sequences capable of driving gene expression, we defined active sequences as those with RNA/DNA ratios above the median of positive controls in their respective libraries (DA: 1.047, variant:1.068), conservatively treating the remaining sequences as inactive in bulk tissue. This definition was highly concordant with MRPAnalyze modeling (25, 43). Combining both libraries, we identified a total of 46,802 active sequences (Fig. 1C) and 25,557 with activity above the 75th percentile of the positive controls. Compared with inactive sequences, active sequences are significantly more conserved (P = 5.8 × 10−28, Wilcoxon test), and their target genes are expressed at higher levels during mid-gestation (P = 6.4 × 10−6, Wilcoxon test; 23% of all sequences mapped to target genes using PLAC-seq data) (27). Comparing sequences with activity in the lower versus upper quartile, we found gene ontology (GO) enrichment for neurodevelopmental terms, such as “nervous system development” (Fig. 1D and fig. S2B), as well as enrichment for transcription factor binding sites (TFBS) for neurodevelopmental gene families such as DLX, LHX, and SOX. We also found enrichment for universal stripe factors (USFs) including EGR1, MAZ, and members of the KLF/SP family (Fig. 1E). USFs colocalize at most promoters and enhancers, increasing chromatin accessibility and residence time for cofactors (44), suggesting that they play a similar role with lentiMPRA reporter constructs integrated into the genome. Together, these results indicate that our active sequences have biological functions in brain development.
Thousands of sequences with cell type–specific chromatin accessibility are active enhancers
Of 46,370 DA sequences passing quality control, 24,218 (52%) were active enhancers in primary cortical cells in our bulk lentiMPRA (data S1), with the percentage active ranging from 43 to 62% across sets of DA sequences predicted to be cell type–specific based on their scATAC-seq profiles (Fig. 2A). Many TFBS show positional enrichment within the scATAC-seq DA sequences with activity in the upper versus lower quartile (Fig. 2B). For example, active sequences tend to have ATOH1, NEUROD2, and TCF4 motifs upstream of the peak summit, whereas ASCL1 and SPI1 motifs are enriched downstream. Repressive sequences, defined as 2784 DA sequences with an activity ratio lower than the 10th percentile of negative controls, showed enrichment for the transcriptional repressors ZEB1 and ZEB2 (fig. S2C and data S1).
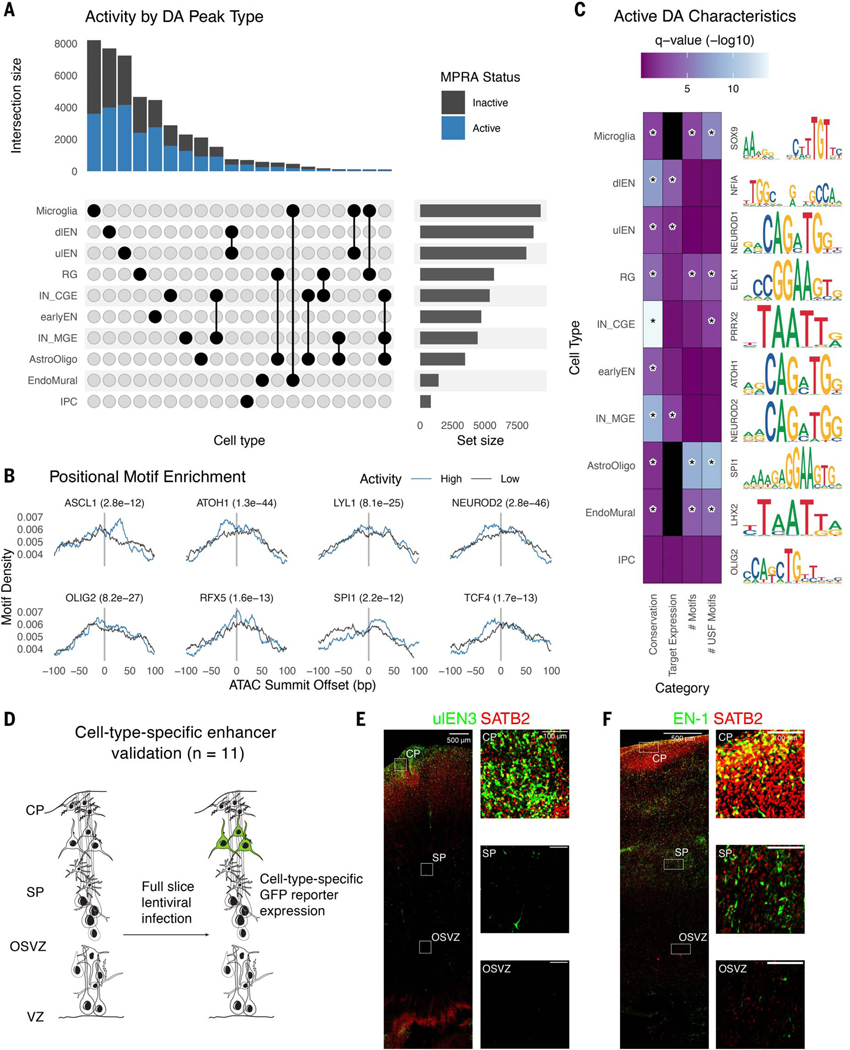
Fig. 2. Identification and validation of functional differentially accessible regions in the developing cortex.
(A) Upset plot showing the number of DA peaks (active, blue; inactive, gray) for each cell type or combination of cell types. (B) The highest activity DA sequences have positional motif enrichment for neurodevelopmental TFs compared with the lowest activity sequences, exhibiting significantly more motif matches slightly up- or downstream of the ATAC-seq peak summit. (C) Active DA sequences have significantly higher means across several attributes compared with inactive DA sequences (color scale is as follows: Wilcoxon test false discovery rate (FDR) adjusted P-values, black indicates no data) including evolutionary conservation (phyloP), expression of PLAC-seq linked target genes in matched cell types, total number of strong motif matches (q-value < 0.01), and total number of strong USF motif matches. A representative motif enriched in active DA peaks for each cell type is shown on the right. Statistically significant comparisons (q-value < 0.05) are indicated by a star. (D) Experimental strategy for validating cell type specificity of active DA sequences. (E and F) Developing human cortex slice cultures transduced with a GFP lentivirus reporter driven by a ulEN-specific enhancer (chr5:89274678–89274948, hg38) (E) and a pan-excitatory neuron specific enhancer (chr2:165141999–165142269, hg38) (F). Expression of GFP (green) and SATB2 (red) was visualized through immunohistochemistry staining and insets show colocalization of GFP+ along with SATB2+ cells in different layers.
In almost all cell types, active DA sequences are more conserved than inactive sequences (Fig. 2C and table S1), consistent with prior knowledge that neurodevelopmental enhancers tend to exhibit strong conservation across vertebrate evolution (45). Inhibitory neurons derived from the ganglionic eminence exhibited the largest differences in conservation scores between active and inactive sequences, fitting with the general transitory role of the ganglionic eminence in guiding neuronal migration (46). To test whether active DAs had regulatory activities endogenously, we predicted the target genes of each DA sequence using PLAC-seq data (27) and calculated cell type– matched expression using scRNA-seq data in the developing human cortex (11). For many neuronal subtypes, genes interacting with active DAs showed higher expression compared with genes interacting with inactive DAs. We also found a higher number of TFBS in active DAs specific to astrocyte/oligodendrocyte precursors (astro/oligo), RG, microglia, and endothelial and mural cells (endo/mural) whereas USF motifs were enriched in IN-CGE DAs and the four glial and vascular cell types. These results indicate that the activity of DA sequences in lentiMPRA is associated with motif content and target gene expression in the matched cell type.
To verify the cell type–specific activity of active DAs in our lentiMPRA, we selected 11 DA peaks with high MPRA activity for six different cell types (table S2) and tested them individually for their enhancer function (Fig. 2D). We infected tissues with the individual enhancer reporter lentivirus and found that all candidates showed GFP expression (fig. S3). Cell-type specificity was inferred from GFP spatial location, counterstaining with cell markers, and morphology. We found three excitatory neuron-specific DA sequences (EN-1, ulEN-2, and dlEN-2) showing enhancer activity in the expected cell type. A ulEN-specific DA region (ulEN-2, chr5:89274678–89274948, hg38, Fig. 2E) drove GFP expression predominantly in the upper areas of the cortical plate and largely co-localized with SATB2, an upper layer excitatory neuron marker. Using PLAC-seq data (27), we found that this region has an EN-specific interaction with the promoter of MEF2C and MEF2C-AS1, known ASD and SCZ genes with EN-specific expression in the developing cortex. The pan-excitatory neuron specific accessible region (EN-1, chr2:165141999–165142269, hg38, Fig. 2F) showed higher GFP signal in the cortical plate (CP) and subplate (SP) compared with the ventricular zone (VZ) and outer subventricular zone, with most of the cells positive for GFP and SATB2 located in the top layer of the CP.
Not all sequences showed enhancer activity within or unique to their predicted cell type. Two regions (ulEN1 and dlEN1) showed GFP expression outside the regions where the expected cell type is enriched. Candidate sequences specific to astro/oligo or RG showed GFP signal around the VZ but also near the CP. These GFP+ cells showed complex morphologies: some matched with the expected cell type(s) whereas others did not (fig. S3). To conclude, we independently validated the enhancer activities of 11 sequences with high lentiMPRA activity, finding all of them to drive GFP expression in cortical cells, with three exhibiting cell type–specificity consistent with scATAC-seq.
To further validate with an orthogonal method, we performed luciferase reporter assays on 24 DA sequences with scATAC-seq chromatin accessibility specific to either EN or microglia spanning a range of MPRA activity, in bulk cortical cells, purified human excitatory neurons, and microglia (fig. S4). We observed a good agreement between luciferase and MPRA in bulk cortical cells (Pearson correlation = 0.63, P = 0.001) and between bulk versus purified cells (Pearson correlation 0.90 and 0.84, for EN and microglia, respectively). These results suggest that bulk reporter assays can accurately capture the regulatory potential of differentially accessible regions.
lentiMPRA identifies functional regulatory variants
In the variant library, 15,335 variants had both alleles passing quality control and 8029 showed enhancer activity from at least one allele. Most of these active variants had modest effects on enhancer activity (median absolute log2 FC = 0.069) (Fig. 3A). At a 10% FDR in our limma differential activity analysis, 164 out of 8029 variants (2.04%) showed significant allelic effects with the number of down-regulating and up-regulating variants being similar (51% versus 49%, Fig. 3B and data S2). This is in line with previous eQTL analyses which find that ~1% of single nucleotide changes are associated with marked changes in gene expression (47). Among these 164 differentially active variants (DAVs), 26 were in LD with GWAS SNPs and 138 were within 100 kb of differentially expressed disease genes, which is similar to our expectations given the library design (17% GWAS and 83% eQTL). Consistent with being QTLs, DAVs are not enriched for low-frequency variants (OR = 0.8, P = 0.34) nor do they have elevated conservation (OR = 0.88, P = 0.52). Separating DAVs based on scATAC-seq cell type showed enrichment in astro/oligo (OR = 2.39, P = 0.14, Fig. 3C and fig. S5A).
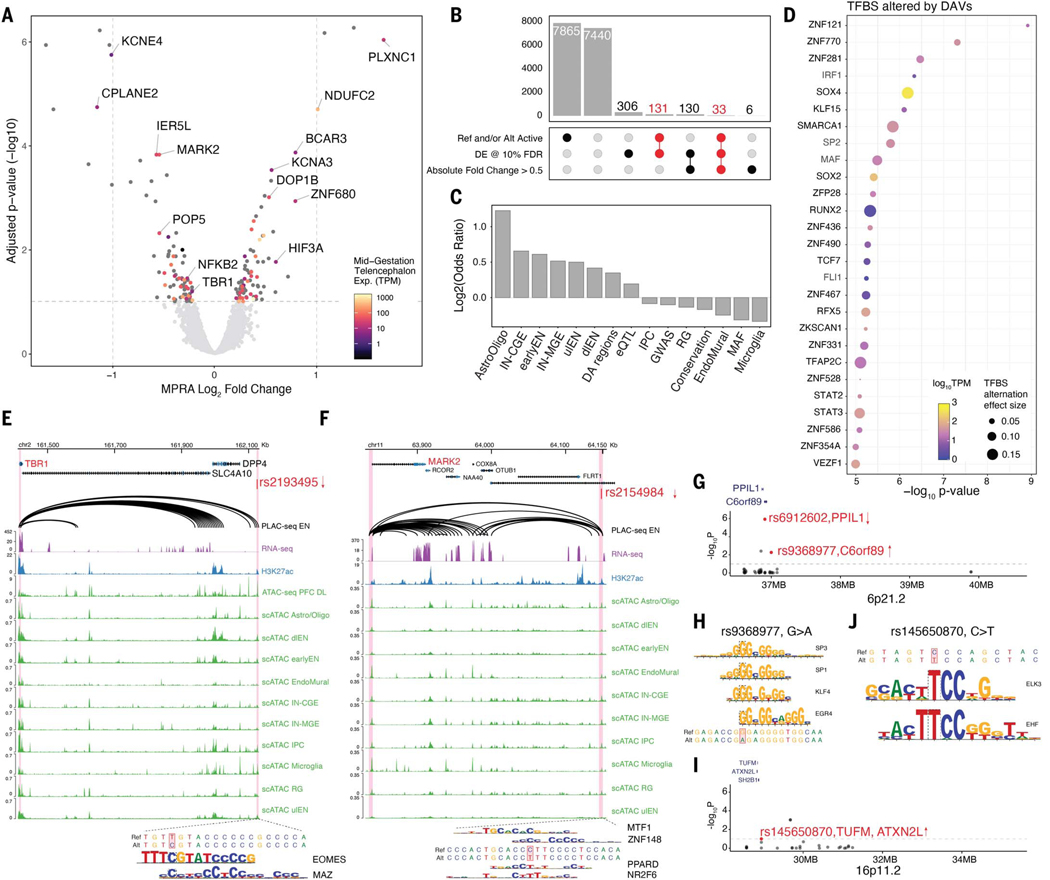
Fig. 3. Identification of differentially active variants associated with psychiatric disorders.
(A) Volcano plot showing log2 fold change and −log10 FDR adjusted P-value for variants that have enhancer activity from at least one allele. Significant variants (FDR < 0.1) were annotated with the PLAC-seq predicted target gene name and color-coded based on target gene expression in mid-gestation telencephalon. Two vertical dashed lines indicate the absolute log2FC of 1. The horizontal dashed line indicates FDR at 10%. (B) Upset plot showing the number of variants (bar) passing combinations of different thresholds (dots and lines below bar). The number of DAVs was highlighted in red. (C) Enrichment log2 odds ratio of DAVs overlapping different features, including combined or separate cell type–specific DA regions, adult brain eQTL, GWAS of various psychiatric disorders and low-frequency variants with minor allele frequency (MAF) less than 0.01. (D) TFBSs predicted to be altered by DAVs using motifbreakR. Dot color represents TF expression in primary cortical cells, size represents predicted magnitude of binding affinity alternation. TFs were ranked by TFBS alternation significance (motifbreakR −log10 P-value, y-axis). (E and F) Genomic browser tracks showing examples of causal regulatory variants and their predicted target genes. The top track shows PLAC-seq chromatin loops in EN (27), the second track shows bulk RNA-seq in primary cortical cells, the third track shows bulk H3K27ac ChIP-seq (26), followed by a track of bulk ATAC-seq in deep-layer cortex (26). The bottom ten tracks show scATAC-seq in the human cortex (11). DAV rs2193495 (E), located in a dlEN-specific accessible region, potentially down-regulates TBR1 expression due to the introduction of EOMES and MAZ binding sites. DAV rs2154984 (F) is predicted to regulate MARK2 expression and disrupt MTF1 and ZNF148 and introduce PPARD and NR2F6 binding sites. (G) Manhattan plot of SCZ-associated chromosome band 6p21.2 showing the 38 variants tested. The y-axis shows −log10 of adjusted P-value from MPRA. DAVs are highlighted in red and annotated with their predicted target gene. Arrows indicate the direction of allele effect observed in MPRA. (H) DAV rs9368977 located in 6p21.2 is predicted to disrupt binding of SP3, SP1, KLF4, and EGR4. (I) Manhattan plot of ASD-associated chromosome band 16p11.2 showing the 25 variants tested. The y-axis shows -log10 of adjusted P-value from MPRA. DAVs are highlighted in red and annotated with predicted target genes. The arrow indicates the direction of allele effect observed in MPRA. (J) TFBS altered in rs145650870. The alternative allele favors the binding of EHF and ELK3.
Next, we compared our DAVs to prior studies. A recent MPRA for dementia-associated variants in human embryonic kidney cells (HEK293T) (14) included 96 variants that were also in our library. We found that 89 variants show no significant allelic effect in either study and 7 altered enhancer activity in HEK293T cells but not in our primary cortical cell data. This difference could be due to the cell types and/or the thresholds used to assign differential activity. Comparing our DAVs to eQTL data from psychENCODE (29), we found that 55% of DAVs (n = 77) had effects in the same direction as the eQTL. The correlation between MPRA and eQTL in non-DAVs (Pearson’s r = 0.008, P = 0.366) was notably lower than that in DAVs (Pearson’sr= 0.14, P = 0.102) (fig. S5B). This corroborates that our lentiMPRA can identify functional variants while underscoring differences between reporter activity and endogenous gene expression.
To decode the mechanisms through which the 164 DAVs exhibit differential activity, we predicted losses and gains of TFBS using motifbreakR (48) (threshold = 1 × 10−5), identifying 34 DAVs (21%) in which the alternative allele alters at least one motif (Fig. 3D). DAVs showed significantly more disruption compared with non-DAVs (OR = 1.49, P = 0.047, Fisher’s exact test). We then analyzed whether these disrupted TFs functionally or physically interact with each other using the STRING database (49) and found a significant TF network centering on SOX2 and STAT3 (PPI enrichment P < 1 × 10−16, fig. S5C).
We predicted the putative target gene/s of DAVs using chromatin interaction data in various brain cell types (27, 28, 50) and adult brain eQTLs (29, 31) (Fig. 3A), finding 48 DAVs (29.3%) to have chromatin loops with gene promoters and 8 of these (17%) to be eQTLs for the interacting gene. As regulatory activities vary over development, target genes predicted using adult brain eQTLs may not reflect genes regulated in early brain development and thus we prioritized target genes predicted from chromatin interaction data. Many target genes are known risk genes or within susceptibility loci for psychiatric disorders and neural diseases. For example, variant rs2193495 is located in a dlEN-specific DA region and potentially regulates the expression of TBR1, a haploinsufficient ASD-associated gene. The rs2193495 alternative allele leads to reduced MPRA activity, possibly due to the creation of EOMES and MAZ binding sites (Fig. 3E). Another down-regulating variant, rs2154984, resides in a putative enhancer of MARK2, a risk gene whose loss-of-function variants have been associated with ASD (51) (Fig. 3F). This variant decreases the binding affinity of MTF1 and ZNF148 while increasing the affinity of PPARD and NR2F6 (Fig. 3F). Another example includes SCZ-associated variant rs10786689 that is thought to regulate NFKB2 and SUFU. This variant decreases enhancer activity, possibly due to the disruption of a SOX2 and/or SOX4 TFBS (fig. S5D). Furthermore, ChIP-seq in human neural progenitor cells (hNPCs) and human embryonic stem cells (hESCs) shows SOX2 binding in this region (52). Because both genes are up-regulated in SCZ patients (53, 54), our results suggest that the rs10786689 alternative allele could be protective. Finally, a down-regulating variant rs73392121 resides within a microglia DA region and is predicted to regulate NPC1, a known cause of Niemann-Pick disease type C. Mutations in this gene lead to impaired cholesterol and lipid cellular transport, including microgliosis (55). Together, these findings demonstrate that lentiMPRA can nominate candidate causal variants for known disease genes.
As a second strategy for linking DAVs to psychiatric disorders, we focused on known risk loci with multiple variants tested in our lentiMPRA. For example, in the SCZ-associated region 6p21.2 (56) (Fig. 3G), we tested 38 variants and found 2 DAVs: rs6912602 and rs9368977. rs6912602 is one of the most differentially active variant in our lentiMPRA (3.3-fold decrease) and is an eQTL associated with reduced expression of PPIL1. Partial loss-of-function variants in PPIL1 cause neurodegenerative pontocerebellar hypoplasia in humans and mice (57). rs9368977 increases enhancer activity and is an eQTL for C6orf89. The alternative allele of rs9368977 disrupts the motifs of USFs SP3 and KLF4 (Fig. 3H). In the SCZ-associated locus 6p21.1 (56), we tested 48 variants and identified one DAV, rs1343025. The alternative allele of rs1343025 is associated with increased expression of VEGFA. VEGFA regulates cerebral blood volume and is associated with SCZ, though the exact impact of VEGFA remains controversial (58). In the ASD risk loci 16p11.2 (59), we tested 25 variants and discovered one activity-increasing DAV, rs145650870 (Fig. 3I). This variant is located in an RG-specific chromatin loop for three nearby genes: TUFM, ATXN2L, and SHSH2B1. The alternative allele of rs145650870 creates a TFBS for EHF (Fig. 3J). Combined, these results show that our lentiMPRA approach could be used to prioritize variants that affect regulatory activity in disease-associated loci.
Organoids show comparable lentiMPRA activity to primary cells
Previous single-cell transcriptomic and epigenomic data along with immunohistochemical analyses suggest that cortical organoids recapitulate many of the cell types in the developing human forebrain (11, 42, 60–62). To explore the MPRA “suitability” of organoids, we tested both our lentiMPRA libraries in 10-week-old cortical organoids (Fig. 4A), validated for the expression of relevant cell type markers through immunostaining (Fig. 4B and fig. S6), bulk RNA-seq (Fig. 4C), and single-cell RNA-seq (fig. S1B). Efficient lentiviral infection in various cell types was also confirmed (fig. S1C). Following 9 weeks of directed differentiation toward a dorsal forebrain fate, organoids were sectioned into 300-mm-thick slices and infected with the lentiMPRA libraries at 10 weeks. This allowed diffusion of lentivirus into most cells, providing high integration rates per cell [multiplicity of infection (MOI) = 100; fig. S7]. Slicing is also known to attenuate hypoxia, leading to better organoid cell health (63). For each library, we infected organoids derived from 2 to 3 iPSC lines with 2 to 4 technical replicates each and analyzed the data as described for primary cells. We observed a high correlation between replicates (average Pearson correlation for DA library: 0.89, variant library: 0.90) and positive controls consistently showed higher enhancer activity compared with negative controls (DA: P = 6.6 × 10−4; variant: P = 0.027, Wilcoxon test; fig. S2A), confirming the high quality of our organoid data.
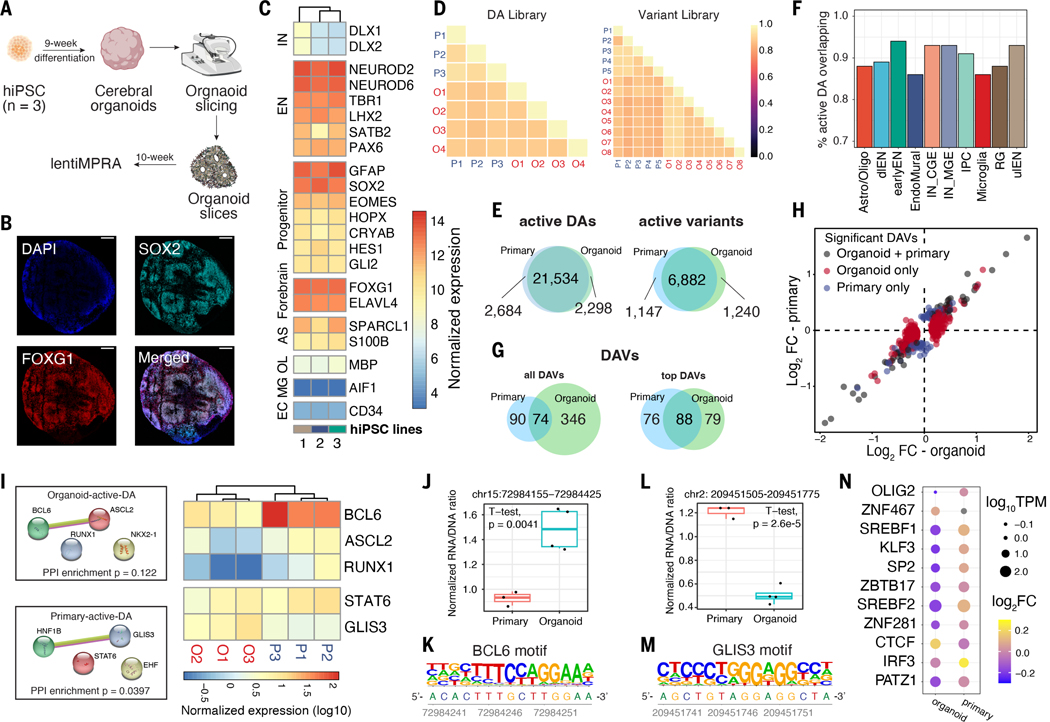
Fig. 4. Comparison of lentiMPRA results in cerebral organoids and primary cortical cells.
(A) Schematic of the experimental workflow. (B) Microscopic images of 10-week-old organoid slices immunostained for SOX2 (Cyan), FOXG1 (Red), and DAPI. Scale bar, 200 μm. (C) Normalized transcript count of marker genes in organoids derived from 3 hiPSC lines (1 = 21792A, 2 = 1323_4, 3 = 20961B). (D) Correlation of log2(RNA/DNA) between replicates in organoids and primary cortical cells for DA library (left) and variant library (right). (E) Venn diagrams showing the overlap between organoids and primary cells. (Left) overlap of active DA regions; (right) overlap of active variants. (F) The proportion of active DAs in organoids that are also active in primary cells. (G) Overlap of DAVs. “Top DAVs” were identified using shuffled sequences to define active and applying a cutoff for absolute log2FC of 0.3. (H) lentiMPRA log2FC in organoid (x-axis) and primary cells (y-axis). The scatter plot includes variants identified as DAVs in both organoids and primary cells (gray), variants detected as DAVs only in organoids (red), and variants detected as DAV only in primary cells (blue). (I) (Left) Protein-protein interactions (PPI) of enriched TFBS motifs in active DAs specific to organoids or primary cells. PPI network generated using STRING (77) database. (Right) heatmap showing the normalized transcript count of enriched TFs from bulk RNA-seq data. TFs not expressed (TPM < 1) in all replicates were removed from the heatmap. (J and K) A DAV (chr15:72984155–72984425, hg38) that contains a BCL6 motif showed increased activity in organoids versus primary cells (J) and its reference sequence contains a BCL6 motif (K). (L and M) A DA region (chr2: 209451505–209451775, hg38) with GLIS3 binding motif showing increased MPRA activity in primary cells versus organoids (L) and the location of the GLIS3 motif in its reference sequence shown below (M). (N) TFBSs altered by DAVs that show an opposite direction of allelic effect between organoids and primary cells. Dot sizes represent normalized TF expression; color represents log2FC.
We compared RNA/DNA ratios between organoids and primary cells and observed high correlation for both libraries (average Pearson correlation DA: 0.89, variant: 0.87, Fig. 4D). Similar to primary cells, roughly half of tested sequences were active (total: 31,954, DA: 23,832, variant: 8122). Most organoid active sequences were also active in primary cells (Fig. 4E). To put this high level of concordance in the context of gene regulation, we performed bulk RNA-seq on three primary and three organoid samples (average replicate Pearson correlation, primary: 0.98, organoid: 0.99) and observed similar transcript levels between primary and organoid samples (average Pearson correlation 0.88), with some notable exceptions discussed below. Finally, we compared the activity of DA sequences stratified by the cell types in which they are accessible and found that active DA sequences were highly concordant in organoids versus primary cells (Fig. 4F). The two cell types having the lowest proportion of primary cell active DAs replicated in organoids were microglia (86.1%) and endothelial cells (86.4%), which is expected as these cell types are thought to be absent in cerebral organoids and our ability to assay activity of these DAs relies upon the permissiveness of MPRAs. These results suggest that cerebral organoids are a reasonable in vitro model of developing forebrain enhancer activity and gene expression, despite some differences in cell type composition and limits to the cell type specificity of bulk MPRAs.
Next, we examined the concordance of differential allelic activity between organoids and primary cells. In organoids, we observed a median absolute log2FC of 0.066, similar to that in primary cells (0.069), and detected 420 DAVs (FDR<10%), of which 74 (18% of organoid DAVs, 45% of primary DAVs) were also DAVs in primary cells (Fig. 4G). The larger number of DAVs identified in organoids is likely due to additional replicates and smaller batch effects. Consistent with this, the overlap of DAV sets was higher (53% of organoid DAVs, 54% of primary DAVs) when considering only the most differentially active organoid variants (absolute log2FC > 0.3 and activity above the median of shuffled controls, Fig. 4G). Despite this modest concordance in which variants were statistically significant, we observed a high correlation in DAV effect sizes in organoids versus primary cells (r = 0.91, P = 2.2 × 10−16, Fig. 4H). We conclude that cerebral organoids and primary cells produce comparable lentiMPRA measurements of differential allelic activity for variants with the largest effects, with noise and cell type differences affecting measurements at and below the significance threshold for identifying DAVs.
We next examined the differences in lentiMPRA results between the two settings. Focusing first on the 2298 DA sequences that were active only in organoids and 2684 only in primary cells, we performed motif enrichment analysis and examined the expression level of enriched TFs (Fig. 4I). Organoid-specific active DA sequences were enriched for binding sites of NKX2.1, RUNX, BCL6, and ASCL2. BCL6 is a transcriptional repressor with significantly lower expression in organoids compared with primary cells (FDR adjusted P-value = 8.8 × 10−7), consistent with our observation that sequences harboring BCL6 motifs tend to have higher lentiMPRA activity in organoids. One such example includes a dlEN-specific accessible region containing a BCL6 motif that had significantly higher enhancer activity in organoids (FDR adjusted P-value =7.91 × 10−6; data S3, Fig. 4, J and K). In addition, overexpression of BCL6 is known to inhibit apoptosis (64) and therefore could reflect elevated cell stress in organoids (65). For the primary-specific active DA peaks, we observed enrichment for GLIS3, STAT6, EHF, and HNF1B motifs. Compared with primary cells, organoids showed higher GLIS3 expression (FDR adjusted P-value = 8.21 × 10−6) and we observed higher lentiMPRA activity in primary cells versus organoids for an astro/oligo and IN-MGE DA region containing a GLIS3 motif, suggesting that it may be functioning as a repressor in these primary-specific active sequences (Fig. 4, L and M). Thus, motif analysis helped us identify TFs whose differential expression between primary cells and organoids is associated with shifts in enhancer activity, suggesting repressor versus activator roles for these TFs and underscoring their importance in regulating neurodevelopment.
Although most variants showed highly comparable effect sizes in organoids and primary cells, we found 61 with an opposite direction of effect. Of these variants, 28 were predicted to alter TFBS motifs and 50% of altered TFs showed differential expression between organoid and primary in bulk RNA-seq (Fig. 4N). For example, rs112049982 increased enhancer activity in primary cells but decreased activity in organoids and was predicted to improve OLIG2 binding affinity, a maker gene for oligodendrocytes (oligo) and oligodendrocyte precursor cells (OPC), which showed significantly lower expression in organoids (FDR adjusted P-value = 8.19 × 10−28), potentially leading to this difference. This also agrees with prior knowledge that oligo and OPC are extremely rare populations in cerebral organoids (11). Together, these results indicate that despite organoids being a suitable in vitro model, differences in the trans-regulating environment should be carefully examined when interpreting lentiMPRA results.
A sequence-based deep learning model of lentiMPRA activity
Our large dataset of lentiMPRA measurements provided an opportunity to characterize the enhancer code in the developing forebrain by modeling enhancer activity and then decoding the model’s understanding of how sequence variants modulate activity. We designed a deep learning regression model that combines a single convolutional layer to learn motif-like sequence features, followed by two recurrent layers to learn the position, spacing, and orientation of motifs (25). Sequences were one-hot encoded into matrices (270bp × 4 nucleotides per sequence), and the mean RNA/DNA ratio across replicates was used as the regression target variable. For each library we trained a model on sequences from all chromosomes except chromosome 3 (used as a validation set to prevent overfitting during training) and chromosome 4 (held out completely for an independent measure of predictive performance). Controls were included in model training. The variant library also included 15,000 sequences that represent a range of expected activity levels as a result of varying epigenetic similarity to validated brain enhancers in the VISTA database (66). On chromosome 4, the DA and variant models achieved 0.82 and 0.78 Pearson correlation, respectively (Fig. 5A; 0.81 and 0.7 Spearman correlation). The most comparable sequence-to-activity model is DeepSTARR (24), trained on fruit fly STARR-seq data (0.68 Pearson correlation for non-housekeeping genes). Though direct comparisons were not possible as a result of vast differences in assay type and dataset quality, our held-out predictive performance suggested that our model was learning relevant sequence features for predicting MPRA activity. It should be noted that the model shares the same limitations as its training data; namely, training on bulk datasets prevents making cell type–specific predictions, and predictions will be most accurate for the brain region and developmental stage of the MPRA.
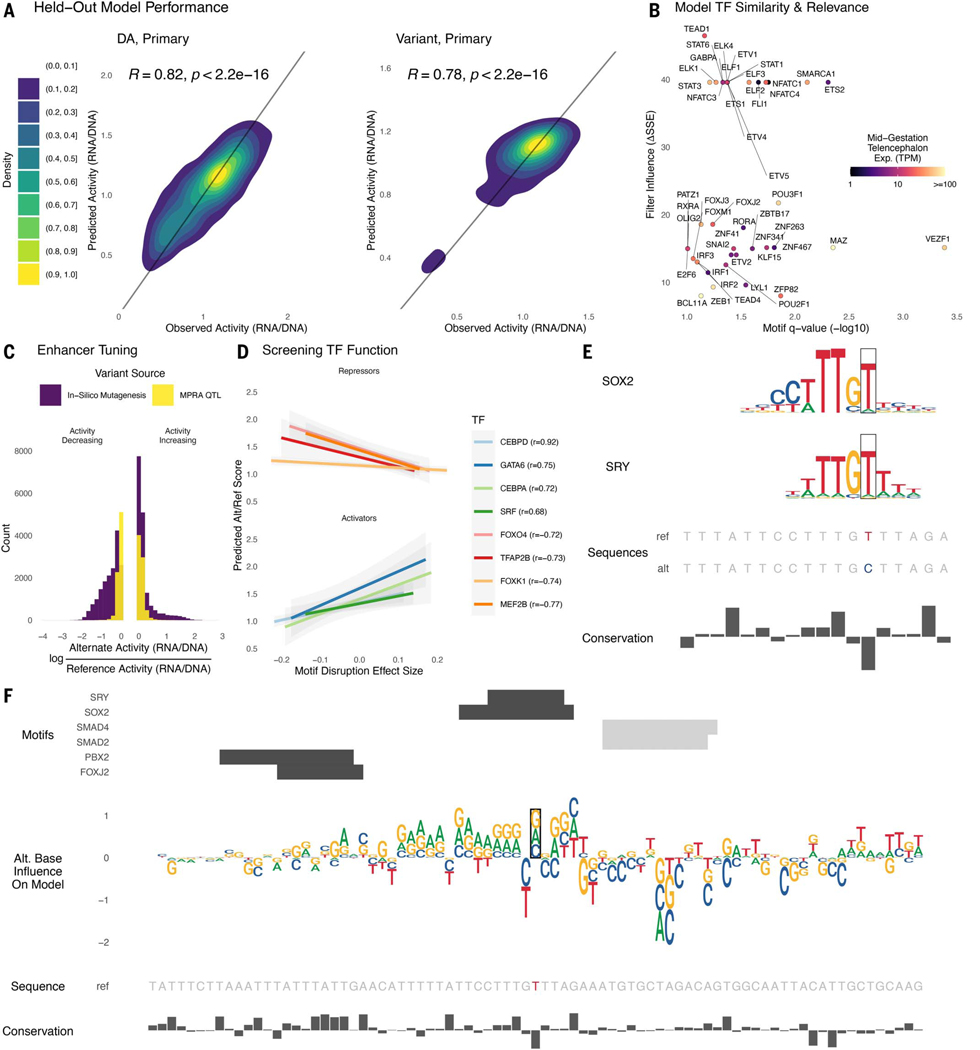
Fig. 5. Sequence determinants of lentiMPRA activity can be modeled with deep learning.
For each library, we trained a deep learning model to predict lentiMPRA activity in primary cells from sequence alone. (A) Sequences on chromosome 4 were held out from model training and used to evaluate model performance. Predicted and measured activity have high Pearson correlation for the DA library (left) and variant library (right). (B) The model learned motifs of neurodevelopmental TFs and used them for accurate predictions. Predictive importance of convolutional filters (change in sum of squared errors when fixing filter output to zero) is plotted against significance of matches to HOCOMOCO motifs (TOMTOM q-value < 0.1) for TFs expressed in developing telencephalon (mean CPM > 1). (C) Applying ISM to the variant library, we found that the activity of most enhancers can be tuned up and down through introduction of alternative alleles. The largest activity-increasing and activity-decreasing alleles for each sequence (purple) tend to have bigger effects than the lentiMPRA measured effects for QTLs (yellow). (D) We combined ISM with motifbreakR TFBS disruption scores to screen TFs for repressor versus activator function in neurodevelopment, using the most activity-changing alternative allele for each sequence in the variant library. TFs where predicted activity is anti-correlated with motif score tend to repress expression (top) and those with a positive correlation tend to be known activators (bottom). This relationship can be used to decode whether the model has learned an activator versus repressor role for TFs that function in both ways. (E) The reference T allele of eQTL rs2883420 (lentiMPRA RNA/DNA 0.8) matches motifs of repressors SRY and SOX2, whereas the alternate C allele disrupts a high information content position in both motifs, resulting in a large activity increase (lentiMPRA RNA/DNA = 0.97, predicted RNA/DNA = 0.96). (F) ISM predicts that the other two possible alleles at rs2883420 also increase activity (middle, sequence logo indicates magnitude and direction: up = increasing, down = decreasing). Alternative alleles at adjacent nucleotides overlapping TF motifs (top, positive strand = black, negative strand = gray) have even larger predicted effects on activity. Region shown is chr10:86,851,230–86,851,500 (hg38).
Convolutional neural networks learn de novo filters from DNA sequences that represent position-specific nucleotide frequencies, similar to TFBS motifs. We therefore used the set of sequences that strongly activate each filter to construct a position weight matrix (PWM) and compared these against the HOCOMOCO (v11) database (67) to identify significant matches to known TFBS (25). As many filters have significant matches to motifs (FDR adjusted P-value < 0.1) for TFs that are expressed in mid-gestation telencephalon (mean TPM > 1), we estimated each filter’s importance for predicting lentiMPRA activity by setting its output to zero and quantifying how much model performance decreased (deltaSSE) (25). Top-ranked filters included TEAD1, NFATC1, STAT3, FOXJ3, POU2F1, and BCL11A (Fig. 5B). In addition to these TFs, our method also highlighted several USFs that function as cofactors to improve chromatin accessibility (44), consistent with our finding that motifs for these TFs are enriched in active versus inactive sequences (Fig. 1E).
To complement this analysis, we performed a large-scale in silico mutagenesis (ISM) study. This method enabled us to quantify how individual nucleotide variants affect model predictions and does not rely upon a PWM database, though we did use PWM similarity to interpret high-scoring variants. Specifically, we constructed sequences with each possible alternate base at each of the 270 positions in each of the 17,069 variant-containing oligos, a total of 18.4 million alleles. We then predicted the activity of each alternate allele. Examining the distribution of the largest predicted activity change (up or down) per oligo, we found that although the QTLs tested in our lentiMPRA generally have moderate activity effects, many of the adjacent synthetic ISM variants have larger effects (Fig. 5C). For 11.6% of oligos, predicted activity can be increased by ≥50% through a single nucleotide change. Conversely, 19.7% of oligos can be reduced by ≤ at least 50% through a single change. As expected, activity-increasing variants frequently create binding sites for transcriptional activators (for example, CEBPD) or mutate binding sites for repressors (for example, FOXK1) that are expressed at mid-gestation whereas activity-decreasing variants do the opposite (Fig. 5, D and E). All sequences contained both increasing and decreasing alleles and in most cases the two variants with the largest absolute ISM scores had opposing effects on activity. At nucleotides with large absolute ISM scores, the three alternative alleles tend to all be increasing or decreasing as expected if the reference base is a high information content position in a TFBS (Fig. 5F).
As an example, we highlight the region around eQTL rs2883420 (Fig. 5, E and F) that has strong matches for SRY-like motifs. ISM predicts that all three alternative alleles at rs2883420 increase activity (predicted RNA/DNA ~0.97). In lentiMPRA the reference allele was inactive (RNA/DNA ~0.8), whereas the alternative allele made the sequence nearly active (RNA/DNA ~0.96), fitting with our prediction. Further examination of the sequence effects of this eQTL (Fig. 5F) found a strong disruption of motifs for repression-capable TFs, such as SOX2 (68) and SRY (69). ISM also predicted increased activity for nonreference alleles in a TFBS-sized region surrounding rs2883420, with most of these having larger effects than the eQTL, consistent with our genome-wide observations (Fig. 5C). These findings indicate that our model is learning de novo PWM-like representations which together form neurodevelopmental regulatory grammar. Such a model can be leveraged to perform ISM, revealing how variants not present in an MPRA alter enhancer activity and TF binding or to design cell type–specific enhancers with precisely tuned activity levels.
Discussion
Gene regulatory elements have a major effect on human brain development and neurodevelopmental disorders. We combined lentiMPRA and deep learning to annotate thousands of regulatory elements in the developing cortex and cerebral organoids. This work provides a large catalog of functional human brain developmental enhancers and variants, along with deep learning models that can accurately predict cell type–specific regulatory regions and variant effects. These functional enhancers and variants will aid in the identification of genetic markers and drug targets, supporting advances in both genetic epidemiology and personalized therapeutics for psychiatric disorders. In addition, it showcases the usability of cerebral organoids for testing regulatory activity in mid-gestation and highlights several differences in the trans-regulating environment that should be taken into account.
One limitation of MPRAs is that they measure the regulatory activity of a sequence but do not identify its target gene. Another caveat is the capability to detect cell type–specific regulatory elements in bulk tissues. This could be due to several factors: (i) Selecting candidate sequences from scATAC-seq, which has low resolution per cell and is descriptive. Nonetheless, nearly half of the 48,861 cell type–specific open chromatin regions that we tested had enhancer potential in primary cells and/or organoids. (ii) Another factor is that MPRA tests sequences outside their native genomic environment. For example, we observed lentiMPRA activity for some microglia- and endothelial-specific DA sequences in organoids, despite these cell types being absent or very rare (11). We hypothesize that this is due to sequences being activated by TFs present in other cell types that do not activate the endogenous sequence because of repressive chromatin. (iii) It is more difficult to detect active regulatory elements for nonabundant cell types in bulk MPRA. We indeed found that abundant cell types, such as neurons and radial glia, had higher percentages of active cell type–specific DA sequences compared with rarer cell populations. For microglia, this could also be due to its resistance to lentivirus infection (70) (fig. S1C), leading to its lower active DA percentage (43.9%). (iv) Our MPRA tested short (270-bp) sequences that could lack additional sequences, which may fine-tune cell type–specificity. (v) Technical differences between our MPRA and the immunostaining and luciferase assays. In addition, as a result of their low throughput nature, only a small number of sequences can be tested. Nonetheless, our validation of 11 regions in developing brain tissues and 24 sequences in sorted EN or microglia cells identified a few sequences showing expected cell-type specificity, whereas the rest were nonspecific. Future studies that utilize single-cell techniques or purified cell populations to validate a larger number of sequences will enable a more comprehensive analysis of the complexity of cell type–specific regulation.
The cerebral organoids produced highly consistent lentiMPRA measurements for the same sequences in primary cortical cells. It is worth noting that the high concordance may, in part, be attributed to the “permissiveness” of MPRA. However, our bulk RNA and scRNA-seq data also showed significant consistency between primary cortex tissue and cortical organoids. Although differential allelic activity was highly correlated for the variants with the largest effects, at least half of the DAVs identified in organoids or primary cells were not statistically significant in the other context with some having opposite allelic effects. However, we also found that these discordant results could shed light on differences in the cellular environment between these two contexts. We identified BCL6 and GLIS3 as TFs whose differential expression in primary cells versus organoids can explain the lentiMPRA’s differential activity. By analyzing whether motifs are positively or negatively correlated with activity, both this analysis and our deep learning-based ISM analysis showed how lentiMPRA data can be used to infer TF function. These computational inferences are needed as many TFs have both repressive and activating functions [e.g., (42, 71–73)].
We evaluated the regulatory effect of 17,069 brain QTLs linked to psychiatric disorders, identifying 164 differentially active variants. This number is in line with other MPRAs that tested the effect of single-nucleotide variants (13, 14) observing relatively small effects of single nucleotide substitutions, especially common alleles, on regulatory activity. Our deep learning model supports this conclusion; predicting that many nucleotide changes in the same regions we tested, including alleles never or rarely seen in people, would show greater differential activity than brain QTLs. In addition, it is worth noting that we observed a modest correlation between MPRA and eQTL effect sizes. This may highlight the need for further functional validation using alternative methods, such as prime editing screens. Another potential caveat is the use of adult instead of developmental brain QTLs, which could be more relevant for neurodevelopmental disorder–associated genes. Additionally, noncoding variants affect different layers of transcriptional regulation than coding variants. MRPAs detect variants affecting enhancer activity or TF binding (74) but not those that modulate genome folding, splicing, or other gene expression aspects. Finally, because about half of the DAVs we detected are in cell-type–specific open chromatin regions, we expect that performing lentiMPRA on mixed cell populations limits the detection of allelic effects that vary across cellular contexts.
Despite detecting only 164 high-confidence DAVs, integrative analysis of our data with publicly available chromatin interaction data linked many of these DAVs to one or more target genes expressed in neurodevelopment. Predicted target genes of many DAVs are known risk genes or within susceptibility loci, such as TBR1 and MARK2 for ASD or NFKB2 and SUFU for SCZ. In particular, for large psychiatric disorder–associated loci, our results for 6p21.1, 6p21.2, and 16p11.2 showcase the utility of lentiMPRA to identify potential disorder–associated regulatory variants in a high-throughput manner. In summary, we nominated several differentially active QTLs as potential causal variants of known disorder genes/loci, paving the way for developing genetic diagnostic and therapeutic tools.
Overall, our work strengthens the utility of using primary cell culture, organoids, MPRAs, and deep learning to investigate regulatory elements and variants involved in human brain development. Future work may consider utilizing an organoid lentiMPRA approach to test libraries from various psychiatric disorder–derived or nonhuman primates iPSCs. Another technological development that could be used to expand upon this study is single-cell MPRA (75, 76). Although currently limited to a small number of sequences, this approach could eventually overcome some limitations we faced testing cell type–specific DAs in a bulk assay. It will also be critical to leverage CRISPR screens to assess the endogenous activity of candidate regulatory sequences, including those validated for activity with MPRAs, although with their own caveats such as the need for high effect sizes on target gene mRNA levels. Deciphering the regulatory code of human brain development will require integration of all these strategies, and the datasets and models generated in this work are a step in that direction.
ACKNOWLEDGMENTS
We thank members of the Ahituv, Nowakowski, Pollen and Pollard labs for assistance with this manuscript. We would like to thank K. White and S. Gaynor for sharing control sets of MPRA. This article was prepared while M. A. Peters was employed at Sage Bionetworks. The opinions expressed in this article are the author’s own and do not reflect the view of the National Institute on Aging, the National Institutes of Health, the Department of Health and Human Services, or the US government.
Funding:
This work was funded in part by the National Institute of Mental Health (NIMH) grant numbers U01MH116438 (to N.A. and K.S.P.); R01MH109907 (to N.A. and K.S.P.); R01MH123179 (K.S.P.), UF1MH130700 (to T.N.); R01NS123263 (to T.N.); DP2MH122400 (to A.A.P.); New York Stem Cell Foundation (to T.N. and A.A.P.); and R01MH125246 (to N.A.); R56MH114911 (to F.M.V.); the National Human Genome Research Institute grant number UM1HG011966 (to N.A.); and Coordination for the Improvement of Higher Education Personnel (CAPES/Brazil)-Finance Code 001. Data were generated as part of the PsychENCODE Consortium, supported by: U01DA048279, U01MH103339, U01MH103340, U01MH103346, U01MH103365, U01MH103392, U01MH116438, U01MH116441, U01MH116442, U01MH116488, U01MH116489, U01MH116492, U01MH122590, U01MH122591, U01MH122592, U01MH122849, U01MH122678, U01MH122681, U01MH116487, U01MH122509, R01MH094714, R01MH105472, R01MH105898, R01MH109677, R01MH109715, R01MH110905, R01MH110920, R01MH110921, R01MH110926, R01MH110927, R01MH110928, R01MH111721, R01MH117291, R01MH117292, R01MH117293, R21MH102791, R21MH103877, R21MH105853, R21MH105881, R21MH109956, R56MH114899, R56MH114901, R56MH114911, R01MH125516, R01MH126459, R01MH129301, R01MH126393, R01MH121521, R01MH116529, R01MH129817, R01MH117406, and P50MH106934, awarded to: A. Abyzov, N.A., S. Akbarian, K. Brennand, A. Chess, G. Cooper, G. Crawford, S. Dracheva, P. Farnham, M. Gandal, M. Gerstein, D. G., F. Goes, J. F. Hallmayer, V. Haroutunian, T. M. Hyde, A. Jaffe, P. Jin, M. Kellis, J. Kleinman, J. A. Knowles, A. Kriegstein, C. Liu, C. E. Mason, K. Martinowich, E. Mukamel, R. Myers, C. Nemeroff, M. Peters, D. Pinto, K.S.P., K. Ressler, P.R., S. Sanders, N. Sestan, P. Sklar, M. P. Snyder, M. State, J. Stein, P. Sullivan, A. E. Urban, F.M.V., S. Warren, D. Weinberger, S. Weissman, Z. Weng, K. White, A. J. Willsey, H. Won, and P. Zandi, respectively.